Code
# Install these packages if you have not already:
# install.packages(c("ipd", "MASS", "broom", "tidyverse", "patchwork"))
library(ipd)
library(MASS)
library(broom)
library(tidyverse)
library(patchwork)Prediction-Based Inference: Methods & Applications
Build intuition for prediction-based inference by simulating data and comparing different methods.
Welcome to this workshop on Prediction-Based (PB) Inference.
In this first module, we use a simple synthetic example to build intuition for how PB inference works and why it is needed. We will:
ipd package,tidy(), glance(), and augment() methods.Throughout the workshop, each exercise includes a solution and short notes to help connect the results back to the main PB inference ideas.
Unit 00 uses synthetic examples on purpose. The goal is to make the PB inference workflow concrete before moving to real applications in later modules.
Using predicted outcomes can improve efficiency, but treating them as if they were the true observed outcomes can bias downstream inference. PB inference methods use a small labeled set to correct that bias while still borrowing information from unlabeled data and predictions.
In this unit, we will:
ipd::ipd() to estimate the association between Y and X1.First, make sure you have the ipd package and a few supporting packages installed:
# Install these packages if you have not already:
# install.packages(c("ipd", "MASS", "broom", "tidyverse", "patchwork"))
library(ipd)
library(MASS)
library(broom)
library(tidyverse)
library(patchwork)Throughout the workshop, we will use reproducible seeds and tidyverse conventions.
Here is a brief overview of the main ipd functions used in this unit.
ipd()ipd() fits IPD estimators for downstream inference with predicted outcomes.
ipd(
formula, # A formula: e.g., Y - f ~ X1 + X2 + ...
method, # Character: one of "chen", "pdc", "postpi_analytic",
# "postpi_boot", "ppi", "ppi_all", "ppi_plusplus", "pspa"
model, # Character: one of "mean", "quantile", "ols", "logistic",
# "poisson"
data, # Data frame containing columns for formula and label
label, # Character: name of the column with set labels ("labeled" and
# "unlabeled")
... # Additional arguments
)Methods currently implemented are:
chen: Chen and Chen Estimator (Gronsbell et al., 2026)pdc: Prediction Decorrelated Inference (Gan et al., 2024)postpi_analytic: Analytic Post-Prediction Inference (Wang et al., 2020).postpi_boot: Bootstrap Post-Prediction Inference (Wang et al., 2020).ppi: Prediction-Powered Inference (Angelopoulos et al., 2023)ppi_plusplus: PPI++ (PPI with Data-Driven Weighting; Angelopoulos et al., 2024)ppi_a: PPI using All Data (Gronsbell et al., 2026)pspa: Assumption-Lean and Data-Adaptive Post-Prediction Inference (Miao et al., 2024)Like many model objects in R, ipd fits work with familiar helper functions:
print() and summary(): display model summaries.tidy(): return a tibble of estimates and standard errors.glance(): return a one-row tibble of model-level metrics.augment(): return the original data with fitted values and residuals.The ipd package provides a function for simulating data based on the approach in Wang et al. (2020). We provide more detail about this function at the end of this model. Here, we will be generating synthetic data following Gronsbell et al. (2026) and Ji et al. (2025) to better illustrate the core principles of PB inference in a more straightforward manner. We provide helper functions below to simulate data from a linear regression model.
#--- Function for the l2 Norm
l2_norm <- function(x, squared = FALSE) {
y <- sum(x^2)
if (!squared) {
y <- sqrt(y)
}
return(y)
}
#--- Function to Generate from the Unit Sphere S^{d-1}
unit_vec <- function(d) {
v <- rnorm(d)
return(v / l2_norm(v))
}
#--- Function to Simulate Data
simple_data_gen <- function(
n, # Number of Samples
p = 2, # Number of Covariates
prop_lab = 0.1, # Proportion of Labeled Observations
total_var = 20, # Total Variance
ratio_var = 1, # Variance Ratio
rho = 0, # Covariate Correlation
sigma_y = 1, # Outcome Error Variance
beta = c(1,1), # Effect Sizes
predset_ind = 1) { # Indices for Covariates Used for Prediction.
#- Covariate Covariance Matrix
sigma_x <- total_var / (ratio_var + 1)
sigma_w <- ratio_var * sigma_x
cov_diag <- sqrt(c(rep(sigma_x, p / 2), rep(sigma_w, p / 2)))
cov_matrix <- (1 - rho) * diag(cov_diag^2) + rho * (cov_diag %o% cov_diag)
#- Generate Covariates
x <- MASS::mvrnorm(n, mu = rep(0, p), Sigma = cov_matrix)
colnames(x) <- paste0('x', 1 : p)
#- Generate Outcome
eps <- rnorm(n, sd = sigma_y)
lin_pred <- x %*% beta
y <- lin_pred + eps
#- Inference Data
inference_data <- data.frame(y = y, x)
#- Generate Predictions
pred_covs <- as.matrix(inference_data[, paste0("x", predset_ind)])
pred_beta <- c(beta[predset_ind])
f <- pred_covs %*% pred_beta
#- Analysis Data
analysis_data <- data.frame(inference_data, f)
samp_ind <- rbinom(n, 1, prop_lab)
analysis_data$set_label <- ifelse(samp_ind == 1, "labeled", "unlabeled")
return(as_tibble(analysis_data))
}Let us generate a synthetic dataset for a linear model with:
n = 25000 samplesp = 10 covariatesprop_lab = 0.1 proportion of labeled observationstotal_var = 20ratio_var = 1rho = 0)sigma_y = 5 outcome error variancebeta <- c(unit_vec(p / 2), unit_vec(p / 2)) generated from the unit circlepredset_ind = 1 : 10 indices for covariates used for predictionThe first setting is an ‘ideal’ setting that mimics when the prediction is derived from a linear model that was estimated on a large, independent dataset with the same distribution of \(Y | X\) as the dataset used for conducting inference. Here, we generate predictions based on all \(X\)’s being available. In this stylized example, we are generating predictions \(f = E[Y | X]\) so the naive approach should be unbiased, but still anti-conservative, as we do not fully capture the variability of \(Y\).
set.seed(123)
n <- 25000 # Number of Samples
p <- 10 # Number of Covariates
b1 <- unit_vec(p / 2) # Effect Sizes
b2 <- unit_vec(p / 2)
dat_ipd <- simple_data_gen(
n = n, # Number of Samples
p = p, # Number of Covariates
prop_lab = 0.1, # Proportion of Labeled Observations
total_var = 20, # Total Variance
ratio_var = 1, # Variance Ratio
rho = 0, # Covariate Correlation
sigma_y = 5, # Outcome Error Variance
beta = c(b1, b2), # Effect Sizes
predset_ind = 1:10 # Indices for covariates used for prediction.
)# The resulting tibble `dat_ipd` has columns:
# - X1, X2, ..., Xp: Simulated covariates
# - Y : True outcome
# - f : Predicted outcome
# - set_label : {"labeled", "unlabeled"}
# Quick look:
dat_ipd |>
group_by(set_label) |>
summarize(n = n()) # A tibble: 2 × 2
set_label n
<chr> <int>
1 labeled 2551
2 unlabeled 22449Let us also inspect the first few rows of each subset:
# Labeled set
dat_ipd |>
filter(set_label == "labeled") |>
glimpse()Rows: 2,551
Columns: 13
$ y <dbl> -10.6651305, 7.0229562, 1.8466360, 5.6049078, 4.4638744, -6.…
$ x1 <dbl> 2.8081938, 1.4447090, 1.1134905, 1.2321898, -0.2950160, 4.69…
$ x2 <dbl> -2.1727482, 1.1195759, 0.1438820, 4.8518833, -1.0137343, 4.4…
$ x3 <dbl> -7.07650307, 0.77112369, -0.01939504, -0.66786699, -4.538479…
$ x4 <dbl> 2.1733859, 3.8373017, 0.3445341, 1.6177814, -1.8449544, -3.7…
$ x5 <dbl> -0.5130663, 2.9121691, 5.2980682, 1.0774526, -1.0392365, -0.…
$ x6 <dbl> -4.32514921, 2.60046203, -0.06035815, -0.23868960, 6.9521900…
$ x7 <dbl> -0.75585286, 3.65797447, 5.16160001, 6.18270859, -4.01256688…
$ x8 <dbl> -3.31953869, 0.07335939, -2.50340399, -4.51835316, -0.851934…
$ x9 <dbl> -1.26218897, -3.11836916, 2.31364284, -0.54576984, -1.158902…
$ x10 <dbl> -1.7577240, -3.5991050, -1.2031549, -1.5884862, 0.9598418, -…
$ f <dbl> -7.981037332, 4.672805866, 1.897680530, 2.421068035, 0.81227…
$ set_label <chr> "labeled", "labeled", "labeled", "labeled", "labeled", "labe…# Unlabeled set
dat_ipd |>
filter(set_label == "unlabeled") |>
glimpse()Rows: 22,449
Columns: 13
$ y <dbl> 0.520872, -7.907675, -6.070025, 6.041134, -3.091775, 5.50057…
$ x1 <dbl> -3.7409669, 3.7360163, -3.2863856, 3.2313674, 3.7157033, 0.1…
$ x2 <dbl> 0.3358049, 3.8872286, -0.2270680, 0.9662234, 1.0364182, -2.6…
$ x3 <dbl> -3.7769611, 0.9883840, -4.4891339, 5.7711764, 0.6166084, 4.0…
$ x4 <dbl> 4.3382493, -4.2897707, 2.5735992, 2.2532995, -5.4164657, -4.…
$ x5 <dbl> -6.12151716, 1.01182690, -1.32007492, 1.62906763, -1.1307161…
$ x6 <dbl> -1.88428832, -2.32594242, 0.22120367, 3.69702327, -0.4563829…
$ x7 <dbl> -0.8790863, 0.1693632, 0.2598034, 1.8054718, 1.1521277, -0.1…
$ x8 <dbl> -5.64360288, -0.41344329, -1.16952884, -0.57129676, 1.196338…
$ x9 <dbl> 0.8406827, 2.6579939, -3.1565336, 4.9834037, 1.5231282, -1.4…
$ x10 <dbl> 3.8708865, 1.1378312, 1.2673506, 0.3500095, 5.6507155, 1.574…
$ f <dbl> -2.0778359, -3.4210568, -1.4953638, 6.2205242, -3.4129538, 6…
$ set_label <chr> "unlabeled", "unlabeled", "unlabeled", "unlabeled", "unlabel…set_label == "labeled" also have both Y and f. In practice, f will be generated by your own prediction model. Here, we do so automatically.set_label == "unlabeled" have Y for posterity (but in a real‐data scenario, you would not know Y). We still generate Y, but the PB routines will not use these. The column f always contains ‘predicted’ values.Before modeling, it is helpful to see graphically how the predicted values, \(f\), compare to the true outcomes, \(Y\).
We can visually assess the bias and variance of our predicted outcomes, \(f\), versus the true outcomes, \(Y\), in our analytic data by plotting:
# Prepare data
dat_visualize <- dat_ipd |>
select(x1, y, f) |>
pivot_longer(y:f, names_to = "Measure", values_to = "Value") |>
arrange(desc(Measure))
# Scatter + trend lines
ggplot(dat_visualize, aes(x = x1, y = Value, color = Measure)) +
theme_minimal() +
geom_point(alpha = 0.4) +
geom_smooth(method = "lm", se = FALSE) +
scale_color_manual(values = c("steelblue", "gray")) +
labs(
x = "X1",
y = "True Y or Predicted f",
color = "Measure"
) +
theme(legend.position = "bottom") 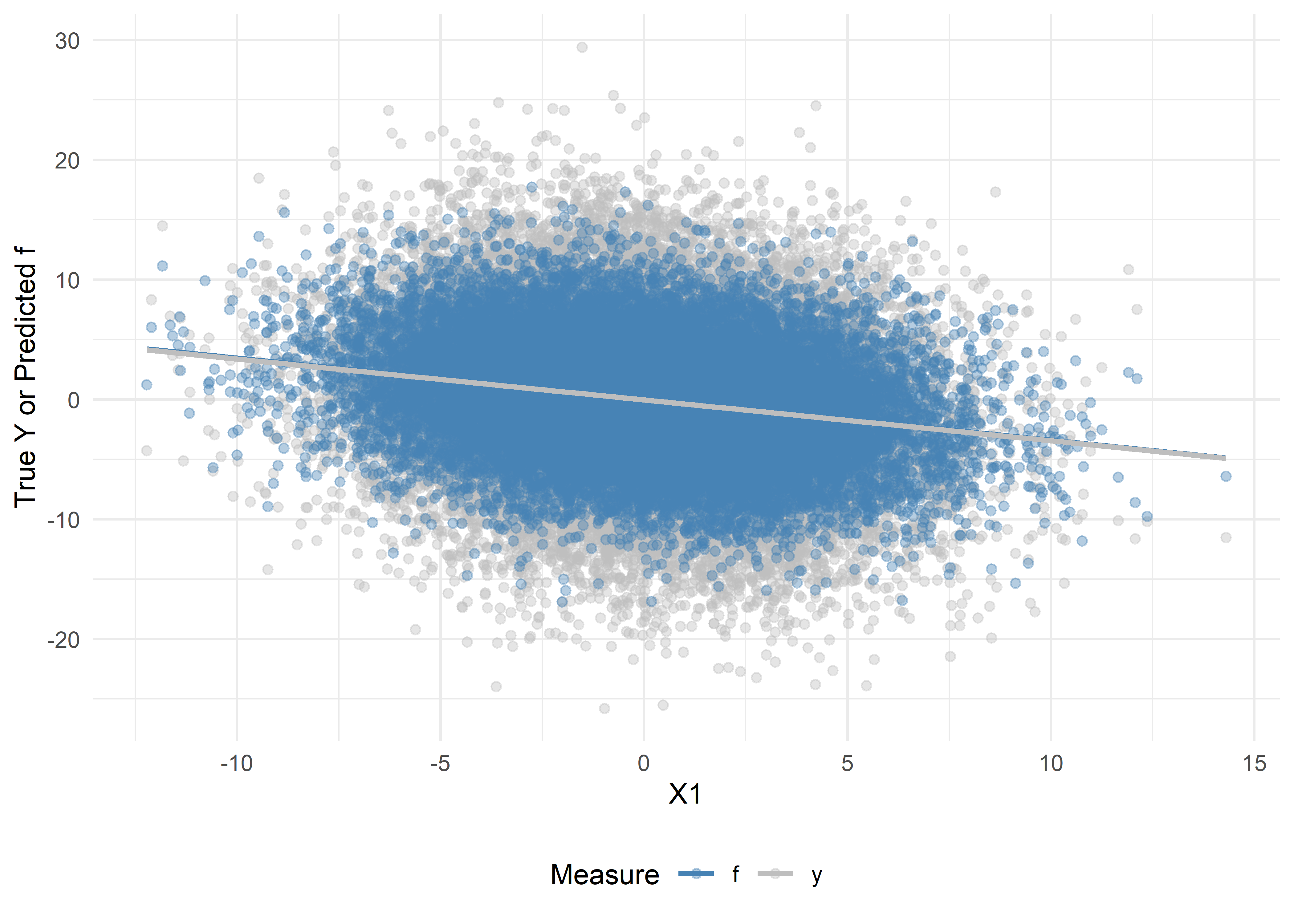
# Density plots
ggplot(dat_visualize, aes(x = Value, fill = Measure)) +
theme_minimal() +
geom_density(alpha = 0.4) +
scale_fill_manual(values = c("steelblue", "gray")) +
labs(
x = "Value",
y = "Density",
fill = "Measure"
) +
theme(legend.position = "bottom")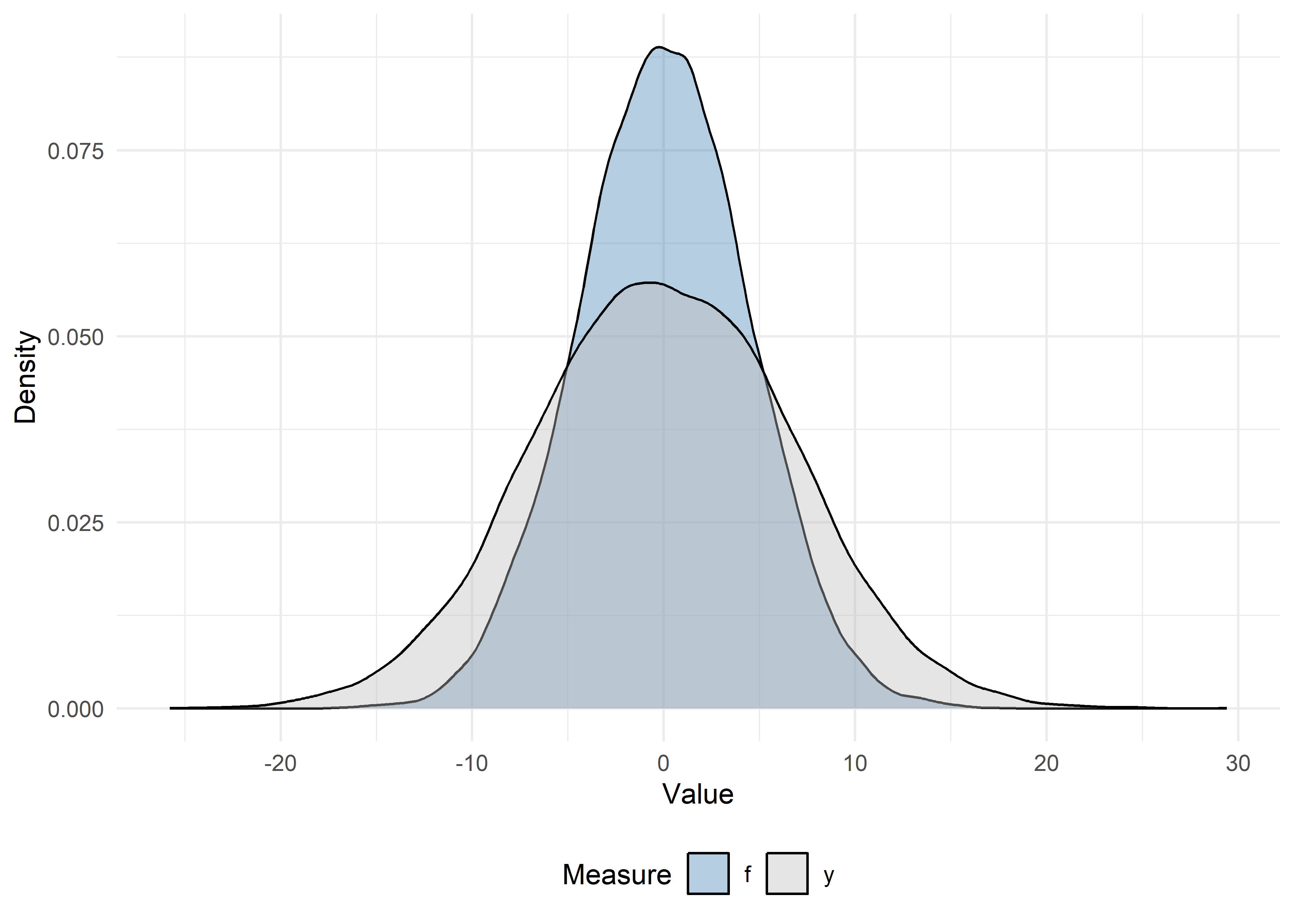
Before applying PB inference, let’s see what happens if we:
Y on X1 for all observations (the oracle approach).f on X1 (the naive approach).Y on X1 (the classical approach).We will compare these to PB inference‐corrected estimates.
Using the labeled and unlabeled sets, fit three models:
lm() with y ~ x1lm() with f ~ x1.lm() with y ~ x1.# 1) Oracle: use the true outcomes for the full data (not possible in practice)
oracle_model <- lm(y ~ x1, data = dat_ipd)
# 2) Naive: treat f as if it were truth (only on unlabeled)
naive_model <- lm(f ~ x1, data = filter(dat_ipd, set_label == "unlabeled"))
# 3) Classical: regress true Y on X1, only on the labeled set
classical_model <- lm(y ~ x1, data = filter(dat_ipd, set_label == "labeled"))Let’s also extract the coefficient summaries using the tidy method and compare the results of the three approaches:
oracle_df <- tidy(oracle_model) |>
mutate(method = "Oracle") |>
filter(term == "x1") |>
select(method, estimate, std.error)
naive_df <- tidy(naive_model) |>
mutate(method = "Naive") |>
filter(term == "x1") |>
select(method, estimate, std.error)
classical_df <- tidy(classical_model) |>
mutate(method = "Classical") |>
filter(term == "x1") |>
select(method, estimate, std.error)
bind_rows(oracle_df, naive_df, classical_df)# A tibble: 3 × 3
method estimate std.error
<chr> <dbl> <dbl>
1 Oracle -0.341 0.0132
2 Naive -0.340 0.00908
3 Classical -0.356 0.0411 ipd::ipd()The single wrapper function ipd() implements multiple PB inference methods (e.g., Chen & Chen, PostPI, PPI, PPI++, PSPA) for various inferential tasks (e.g., mean and quantile estimation, ols, logistic, and Poisson regression).
Basic usage of ipd():
ipd(
formula = Y - f ~ X1, # The downstream inferential model
method = "pspa", # The PB inference method to run
model = "ols", # The type of inferential model
data = dat_ipd, # A data.frame with columns:
# - set_label: {"labeled", "unlabeled"}
# - Y: true outcomes (for labeled data)
# - f: predicted outcomes
# - X covariates (here X1, X2, X3, X4)
label = "set_label" # Column name indicating "labeled"/"unlabeled"
)Let’s run one method, chen, proposed by Gronsbell et al. (2026). The Chen + Chen estimator is a PB method that combines information from:
Rather than treating the predicted outcomes with the same importance as the true outcomes, the method estimates a data-driven weight, \(\hat{\omega}\), and applies it to the predicted outcome contributions:
\[ \hat{\beta}_\text{cc} = \hat{\beta}_\text{classical} - \hat{\omega}\cdot (\hat{\gamma}_\text{naive}^l - \hat{\gamma}_\text{naive}^\text{all}), \]
where \(\hat{\beta}_{\rm classical}\) is the estimate from the classical regression, \(\hat{\gamma}_{\rm naive}^l\) is the estimate from the naive regression on the labeled data, \(\hat{\gamma}_{\rm naive}^\text{all}\) is the estimate from the naive regression on all the data, and \(\hat{\omega}\) reflects the amount of additional information carried by the predictions. By adaptively weighting the unlabeled information, the Chen + Chen estimator achieves greater precision than by using the labeled data alone, without sacrificing validity, even when the predictions are imperfect.
Let’s call the method using the ipd() function and collect the estimate for the slope of X1 in a linear regression (model = "ols"):
set.seed(123)
ipd_model <- ipd(
formula = y - f ~ x1,
data = dat_ipd,
label = "set_label",
method = "chen",
model = "ols"
)
ipd_modelIPD inference summary
Method: chen
Model: ols
Formula: y - f ~ x1
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -0.224224 0.104333 -2.1491 0.03162 *
x1 -0.330246 0.032247 -10.2412 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1The ipd_model is an S4 object with slots for things like the coefficient, se, ci, coefTable, fit, formula, data_l, data_u, method, model, and intercept. We can extract the coefficient table using ipd’s tidy helper and compare with the naive and classical methods:
# Extract the coefficient estimates
ipd_df <- tidy(ipd_model) |>
mutate(method = "Chen + Chen") |>
filter(term == "x1") |>
select(method, estimate, std.error)
# Combine with oracle, naive, and classical:
compare_tab <- bind_rows(oracle_df, naive_df, classical_df, ipd_df)
compare_tab# A tibble: 4 × 3
method estimate std.error
<chr> <dbl> <dbl>
1 Oracle -0.341 0.0132
2 Naive -0.340 0.00908
3 Classical -0.356 0.0411
4 Chen + Chen -0.330 0.0322 Let’s plot the coefficient estimates and 95% CIs for each of the naive, classical, and PB methods:
# Forest plot of estimates and 95% confidence intervals
compare_tab |>
mutate(
lower = estimate - 1.96 * std.error,
upper = estimate + 1.96 * std.error
) |>
ggplot(aes(x = estimate, y = method)) +
geom_point(size = 2) +
geom_errorbarh(aes(xmin = lower, xmax = upper), height = 0.2) +
geom_vline(xintercept = b1[1], linetype = "dashed", color = "red") +
labs(
title = "Comparison of slope estimates \u00B1 1.96·SE",
x = expression(hat(beta)[X1]),
y = ""
) +
theme_minimal()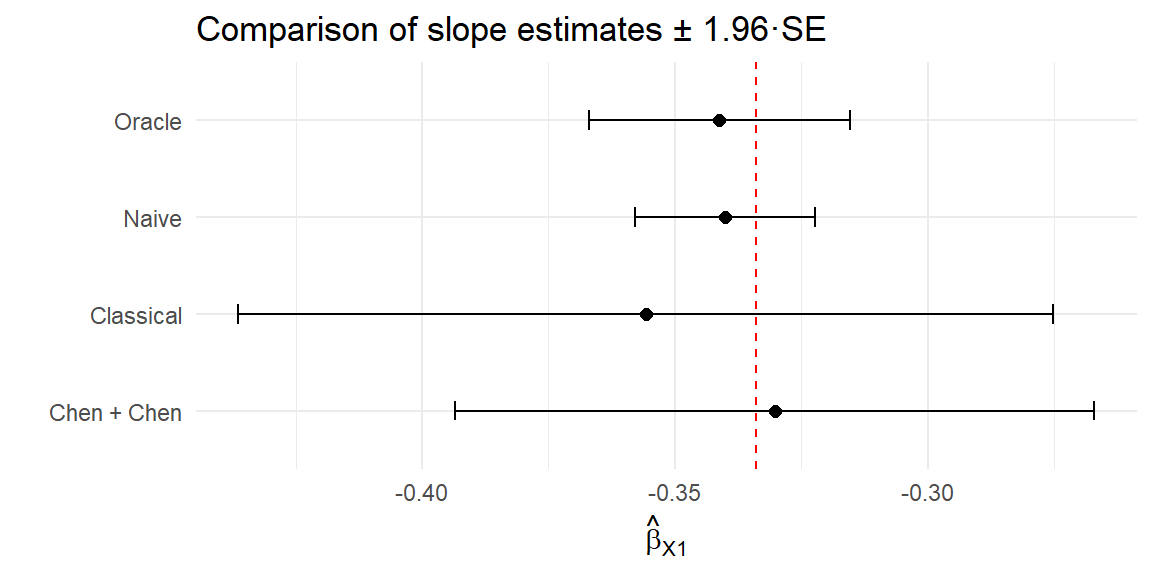
Now, let us repeat Exercises 1-5, but with \(X_1, \ldots, X_5\) omitted from the training of the prediction function:
predset_ind = 6 : 10
We will compare these results with the ideal case in the previous exercises. Here, since the predictions do not include \(X_1\), we should not be able to recover the true association between \(Y\) and \(X_1\).
set.seed(123)
dat_ipd2 <- simple_data_gen(
n = n,
p = p,
prop_lab = 0.1,
total_var = 20,
ratio_var = 1,
rho = 0,
sigma_y = 1,
beta = c(b1, b2),
predset_ind = 6:10 # Omitting X1, ..., X5
)# Prepare data
dat_visualize2 <- dat_ipd2 |>
select(x1, y, f) |>
pivot_longer(y:f, names_to = "Measure", values_to = "Value") |>
arrange(Measure)
# Scatter + trend lines
ggplot(dat_visualize2, aes(x = x1, y = Value, color = Measure)) +
theme_minimal() +
geom_point(alpha = 0.4) +
geom_smooth(method = "lm", se = FALSE) +
scale_color_manual(values = c("steelblue", "gray")) +
labs(
x = "X1",
y = "True Y or Predicted f",
color = "Measure"
) +
theme(legend.position = "bottom") 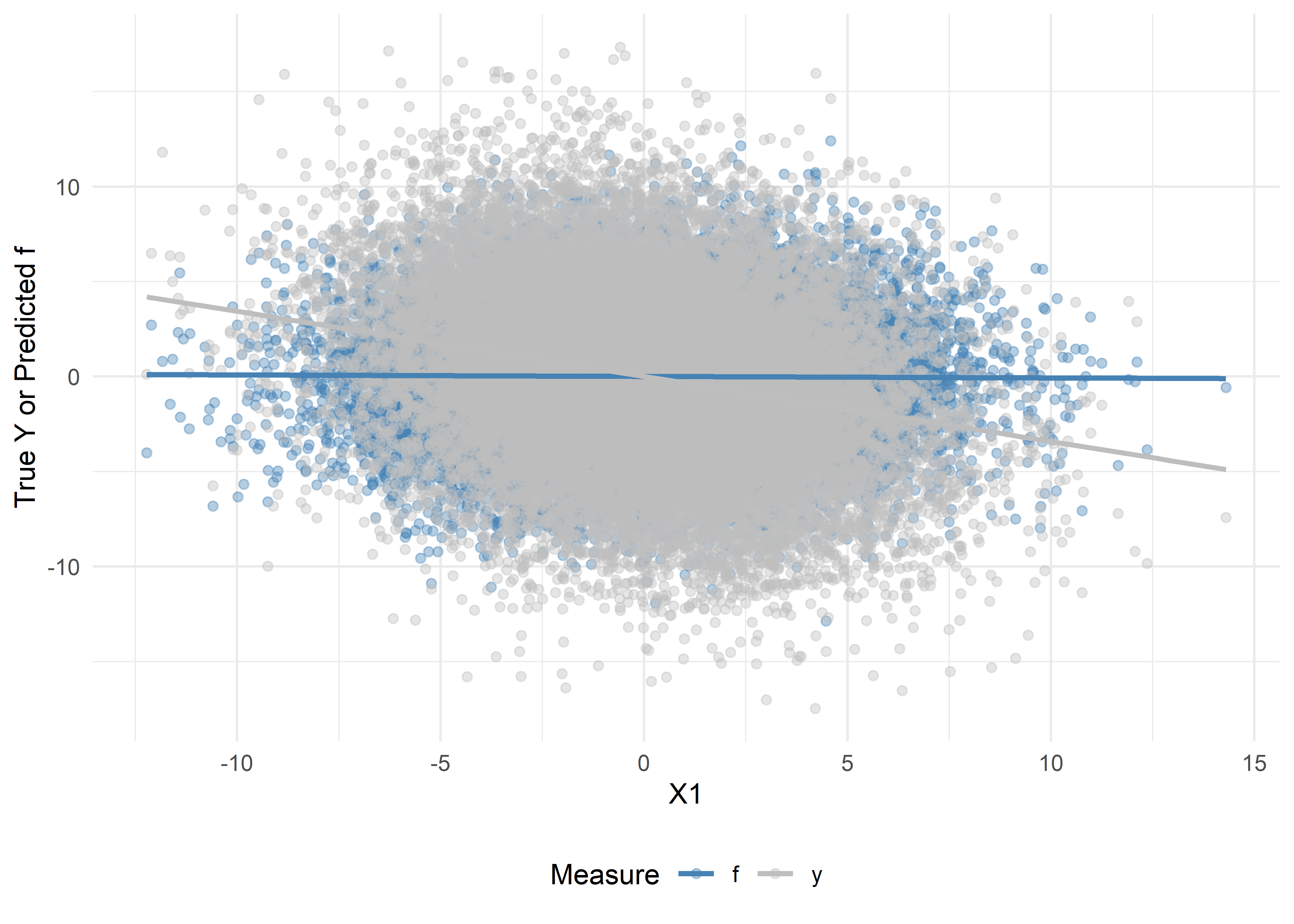
# Density plots
ggplot(dat_visualize2, aes(x = Value, fill = Measure)) +
theme_minimal() +
geom_density(alpha = 0.4) +
scale_fill_manual(values = c("steelblue", "gray")) +
labs(
x = "Value",
y = "Density",
fill = "Measure"
) +
theme(legend.position = "bottom")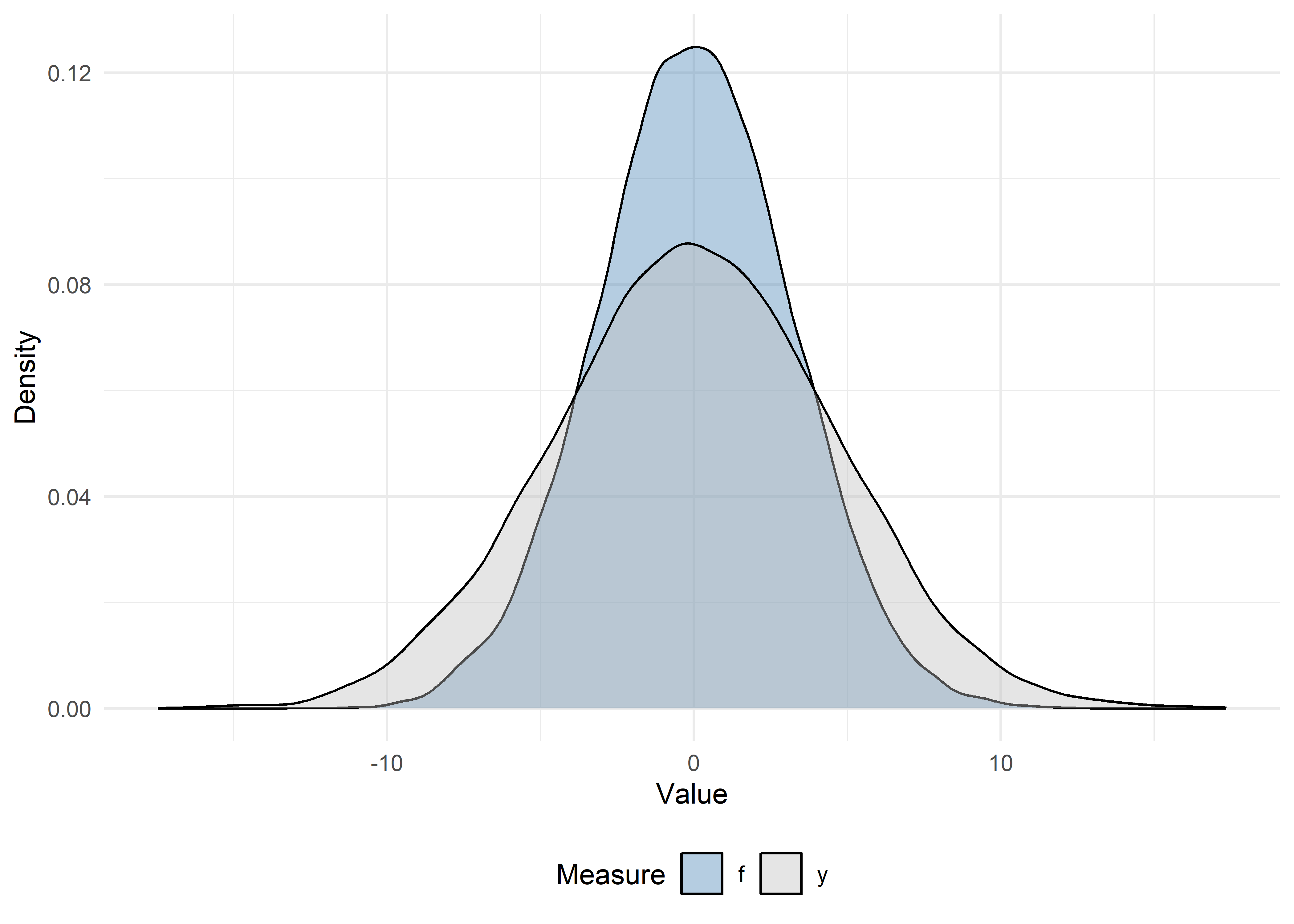
# 1) Oracle: use the true outcomes for the full data (not possible in practice)
oracle_model2 <- lm(y ~ x1, data = dat_ipd2)
# 2) Naive: treat f as if it were truth (only on unlabeled)
naive_model2 <- lm(f ~ x1, data = filter(dat_ipd2, set_label == "unlabeled"))
# 3) Classical: regress true Y on X1, only on the labeled set
classical_model2 <- lm(y ~ x1, data = filter(dat_ipd2, set_label == "labeled"))oracle_df2 <- tidy(oracle_model2) |>
mutate(method = "Oracle") |>
filter(term == "x1") |>
select(method, estimate, std.error)
naive_df2 <- tidy(naive_model2) |>
mutate(method = "Naive") |>
filter(term == "x1") |>
select(method, estimate, std.error)
classical_df2 <- tidy(classical_model2) |>
mutate(method = "Classical") |>
filter(term == "x1") |>
select(method, estimate, std.error)
bind_rows(oracle_df2, naive_df2, classical_df2)# A tibble: 3 × 3
method estimate std.error
<chr> <dbl> <dbl>
1 Oracle -0.343 0.00882
2 Naive -0.00775 0.00664
3 Classical -0.371 0.0272 set.seed(123)
ipd_model2 <- ipd(
formula = y - f ~ x1,
data = dat_ipd2,
label = "set_label",
method = "chen",
model = "ols"
)
ipd_model2IPD inference summary
Method: chen
Model: ols
Formula: y - f ~ x1
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.096086 0.065442 1.4683 0.142
x1 -0.370577 0.020917 -17.7163 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# Extract the coefficient estimates
ipd_df2 <- tidy(ipd_model2) |>
mutate(method = "Chen + Chen") |>
filter(term == "x1") |>
select(method, estimate, std.error)
# Combine with naive & classical:
compare_tab2 <- bind_rows(oracle_df2, naive_df2, classical_df2, ipd_df2)
compare_tab2# A tibble: 4 × 3
method estimate std.error
<chr> <dbl> <dbl>
1 Oracle -0.343 0.00882
2 Naive -0.00775 0.00664
3 Classical -0.371 0.0272
4 Chen + Chen -0.371 0.0209 # Forest plot of estimates and 95% confidence intervals
compare_tab2 |>
mutate(
lower = estimate - 1.96 * std.error,
upper = estimate + 1.96 * std.error
) |>
ggplot(aes(x = estimate, y = method)) +
geom_point(size = 2) +
geom_errorbarh(aes(xmin = lower, xmax = upper), height = 0.2) +
geom_vline(xintercept = b1[1], linetype = "dashed", color = "red") +
labs(
title = "Comparison of slope estimates \u00B1 1.96·SE",
x = expression(hat(beta)[X1]),
y = ""
) +
theme_minimal()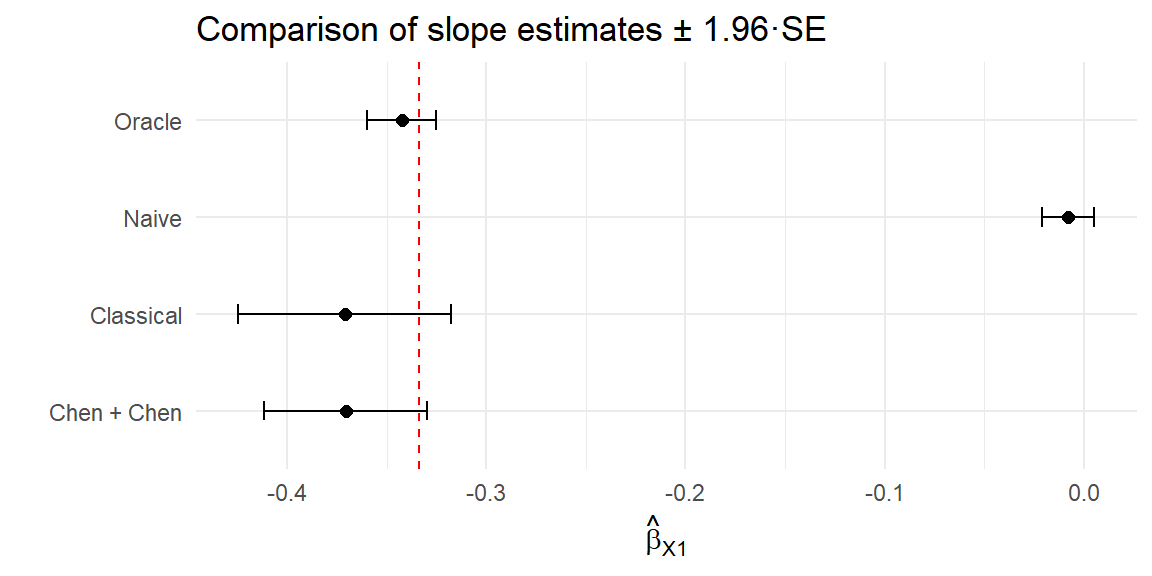
Use tidy(), glance(), and augment() on ipd_model. Compare the coefficient estimate and standard error for X1 with the naive fit.
tidy(ipd_model)# A tibble: 2 × 5
term estimate std.error conf.low conf.high
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -0.224 0.104 -0.429 -0.0197
2 x1 -0.330 0.0322 -0.393 -0.267 glance(ipd_model)# A tibble: 1 × 6
method model intercept nobs_labeled nobs_unlabeled call
<chr> <chr> <lgl> <int> <int> <chr>
1 chen ols TRUE 2551 22449 y - f ~ x1augment(ipd_model) |> glimpse()Rows: 22,449
Columns: 15
$ y <dbl> 0.520872, -7.907675, -6.070025, 6.041134, -3.091775, 5.50057…
$ x1 <dbl> -3.7409669, 3.7360163, -3.2863856, 3.2313674, 3.7157033, 0.1…
$ x2 <dbl> 0.3358049, 3.8872286, -0.2270680, 0.9662234, 1.0364182, -2.6…
$ x3 <dbl> -3.7769611, 0.9883840, -4.4891339, 5.7711764, 0.6166084, 4.0…
$ x4 <dbl> 4.3382493, -4.2897707, 2.5735992, 2.2532995, -5.4164657, -4.…
$ x5 <dbl> -6.12151716, 1.01182690, -1.32007492, 1.62906763, -1.1307161…
$ x6 <dbl> -1.88428832, -2.32594242, 0.22120367, 3.69702327, -0.4563829…
$ x7 <dbl> -0.8790863, 0.1693632, 0.2598034, 1.8054718, 1.1521277, -0.1…
$ x8 <dbl> -5.64360288, -0.41344329, -1.16952884, -0.57129676, 1.196338…
$ x9 <dbl> 0.8406827, 2.6579939, -3.1565336, 4.9834037, 1.5231282, -1.4…
$ x10 <dbl> 3.8708865, 1.1378312, 1.2673506, 0.3500095, 5.6507155, 1.574…
$ f <dbl> -2.0778359, -3.4210568, -1.4953638, 6.2205242, -3.4129538, 6…
$ set_label <chr> "unlabeled", "unlabeled", "unlabeled", "unlabeled", "unlabel…
$ .fitted <dbl> 1.01121686, -1.45803011, 0.86109300, -1.29137161, -1.4513217…
$ .resid <dbl> -0.4903448, -6.4496451, -6.9311184, 7.3325060, -1.6404531, 5…# Compare with naive
broom::tidy(naive_model)# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 0.0215 0.0288 0.745 4.56e- 1
2 x1 -0.340 0.00908 -37.4 1.06e-297"ppi_plusplus"). How do the results compare?model = "logistic" in both simdat() and ipd().ipd::simdat()simdat() generates synthetic datasets for several inferential settings.
simdat(
n, # Numeric vector of length 3: c(n_train, n_labeled, n_unlabeled)
effect, # Numeric: true effect size for simulation
sigma_Y, # Numeric: residual standard deviation
model, # Character: one of "mean", "quantile", "ols", "logistic", "poisson"
... # Additional arguments
)This function returns a data.frame with columns:
X1, X2, ...: covariatesY: true outcome (for training, labeled, and unlabeled subsets)f: predictions from the model (for labeled and unlabeled subsets)set_label: character indicating “training”, “labeled”, or “unlabeled”.The ipd::simdat() function makes it easy to generate:
We supply the sample sizes, n = c(n_train, n_label, n_unlabel), an effect size (effect), residual standard deviation (sigma_Y; i.e., how much random noise is in the data), and a model type ("ols", "logistic", etc.). In this tutorial, we focus on a continuous outcome generated from a linear regression model ("ols"). We can also optionally shift and scale the predictions (via the shift and scale arguments) to control how the predicted outcomes relate to their true underlying counterparts.
simdat()Try generating data with the simdat() function. Let us generate a synthetic dataset for a linear model with:
set.seed(123)
# n_t = 5000, n_l = 500, n_u = 1500
n <- c(5000, 500, 1500)
# Effect size = 1.5, noise sd = 3, model = "ols" (ordinary least squares)
# We also shift the mean of the predictions by 1 and scale their values by 2
dat <- simdat(
n = n,
effect = 1.5,
sigma_Y = 3,
model = "ols",
shift = 1,
scale = 2
)# The resulting data.frame `dat` has columns:
# - X1, X2, X3, X4: Four simulated covariates (all numeric ~ N(0,1))
# - Y : True outcome (available in unlabeled set for simulation)
# - f : Predicted outcome (Generated internally in simdat)
# - set_label : {"training", "labeled", "unlabeled"}
# Quick look:
dat |>
group_by(set_label) |>
summarize(n = n()) # A tibble: 3 × 2
set_label n
<chr> <int>
1 labeled 500
2 training 5000
3 unlabeled 1500Let us also inspect the first few rows of each subset:
# Training set
dat |>
filter(set_label == "training") |>
glimpse()Rows: 5,000
Columns: 7
$ X1 <dbl> -0.56047565, -0.23017749, 1.55870831, 0.07050839, 0.12928774…
$ X2 <dbl> -1.61803670, 0.37918115, 1.90225048, 0.60187427, 1.73234970,…
$ X3 <dbl> -0.91006117, 0.28066267, -1.03567040, 0.27304874, 0.53779815…
$ X4 <dbl> -1.119992047, -1.015819127, 1.258052722, -1.001231731, -0.40…
$ Y <dbl> 3.8625325, -1.6575634, 4.1872914, -3.3624963, 6.9978916, 1.5…
$ f <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, …
$ set_label <chr> "training", "training", "training", "training", "training", …# Labeled set
dat |>
filter(set_label == "labeled") |>
glimpse()Rows: 500
Columns: 7
$ X1 <dbl> -0.4941739, 1.1275935, -1.1469495, 1.4810186, 0.9161912, 0.3…
$ X2 <dbl> -0.15062996, 0.80094056, -1.18671785, 0.43063636, 0.21674709…
$ X3 <dbl> 2.0279109, -1.4947497, -1.5729492, -0.3002123, -0.7643735, -…
$ X4 <dbl> 0.53495620, 0.36182362, -1.89096604, -1.40631763, -0.4019282…
$ Y <dbl> 2.71822922, 1.72133689, 0.86081066, 3.77173123, -2.77191549,…
$ f <dbl> 0.67556303, -0.13706321, -1.75579589, 0.84146158, 0.15512973…
$ set_label <chr> "labeled", "labeled", "labeled", "labeled", "labeled", "labe…# Unlabeled set
dat |>
filter(set_label == "unlabeled") |>
glimpse()Rows: 1,500
Columns: 7
$ X1 <dbl> -1.35723063, -1.29269781, -1.51720731, 0.85917603, -1.214617…
$ X2 <dbl> 0.01014789, 1.56213812, 0.41284605, -1.18886219, 0.71454993,…
$ X3 <dbl> -1.42521509, 1.73298966, 1.66085181, -0.85343610, 0.26905593…
$ X4 <dbl> -1.1645365, -0.2522693, -1.3945975, 0.3959429, 0.5980741, 1.…
$ Y <dbl> -3.8296223, -3.0350282, -5.3736718, 2.6076634, -4.7813463, -…
$ f <dbl> -1.9014253, -0.1679210, -0.3030749, 0.1249168, -0.8998546, -…
$ set_label <chr> "unlabeled", "unlabeled", "unlabeled", "unlabeled", "unlabel…set_label == "training" form an internal training set. Here, Y is observed, but f is NA, as we learn the prediction rule in this set.set_label == "labeled" also have both Y and f. In practice, f will be generated by your own prediction model; for simulation, simdat does so automatically.set_label == "unlabeled" have Y for posterity (but in a real‐data scenario, you would not know Y); simdat still generates Y, but the PB routines will not use these. The column f always contains ‘predicted’ values.In practice, we would take the training portion and fit an AI/ML model to predict \(Y\) from \((X_1, X_2, X_3, X_4)\). This is done automatically by the simdat function, but this optional activity gives you a hands-on implementation.
Fit a linear prediction model on the training data:
# 1) Subset training set
dat_train <- dat |>
filter(set_label == "training")
# 2) Fit a linear model: Y ~ X1 + X2 + X3 + X4
lm_pred <- lm(Y ~ X1 + X2 + X3 + X4, data = dat_train)
# 3) Prepare a full-length vector of NA
dat$f_pred <- NA_real_
# 4) Identify the rows to predict (all non-training rows)
idx_analytic <- dat$set_label != "training"
# 5) Generate predictions just once on that subset (shifted and scaled to match)
pred_vals <- (predict(lm_pred, newdata = dat[idx_analytic, ]) - 1) / 2
# 6) Insert them back into the full data frame
dat$f_pred[idx_analytic] <- pred_vals
# 7) Verify: `f_pred` is equal to `f` for the labeled and unlabeled data
dat |>
select(set_label, Y, f, f_pred) |>
filter(set_label != "training") |>
glimpse()Rows: 2,000
Columns: 4
$ set_label <chr> "labeled", "labeled", "labeled", "labeled", "labeled", "labe…
$ Y <dbl> 2.71822922, 1.72133689, 0.86081066, 3.77173123, -2.77191549,…
$ f <dbl> 0.67556303, -0.13706321, -1.75579589, 0.84146158, 0.15512973…
$ f_pred <dbl> 0.67556303, -0.13706321, -1.75579589, 0.84146158, 0.15512973…lm(Y ~ X1 + X2 + X3 + X4, data = dat_train) fits an ordinary least squares (OLS) regression on the training subset.predict(lm_pred, newdata = .) generates a new f (stored as f_pred) for each row outside of the training set.ranger::ranger()), gradients (xgboost::xgboost()), or any other ML algorithm; PB methods only require that you supply a vector of predictions, f, in your data.Happy coding! Feel free to modify and extend these exercises for your own data.
This is the end of the module. We hope this was informative! For question/concerns/suggestions, please reach out to ssalerno@fredhutch.org